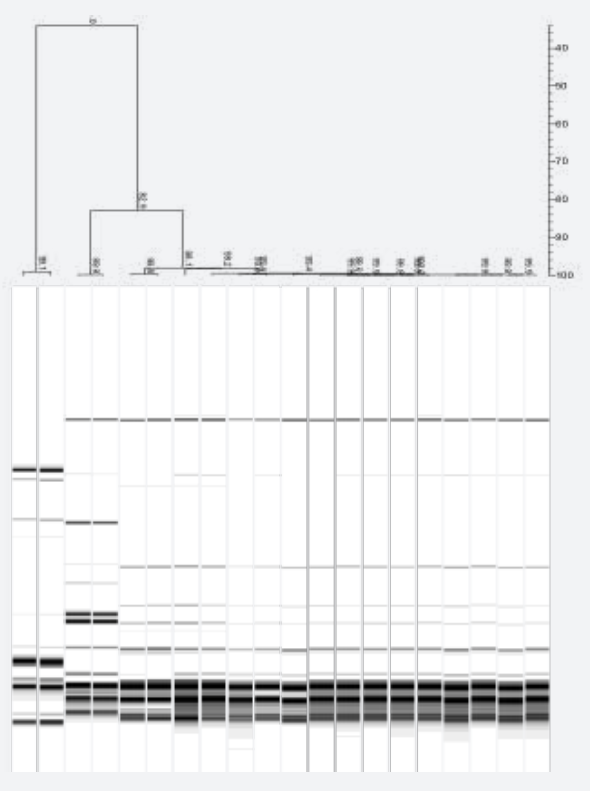
Rep-PCR is a general method applicable for almost all microorganisms. This method generates a DNA fingerprint that is unique to the organism at the strain level. For this we use a combination of GTG5, ERIC1, ERIC2, BOX-A1-R and BOX-A2-R PCR protocols. With a comprehensive, dynamic picture of any microbial environment, technicians can track and manage data at the strain level, letting them pinpoint contamination with a high level of speed, ease and precision. In addition, we offer GMP Microbial strain typing services through our GMP brand ProBase Pharma.
Strain typing using Whole Genome Sequencing
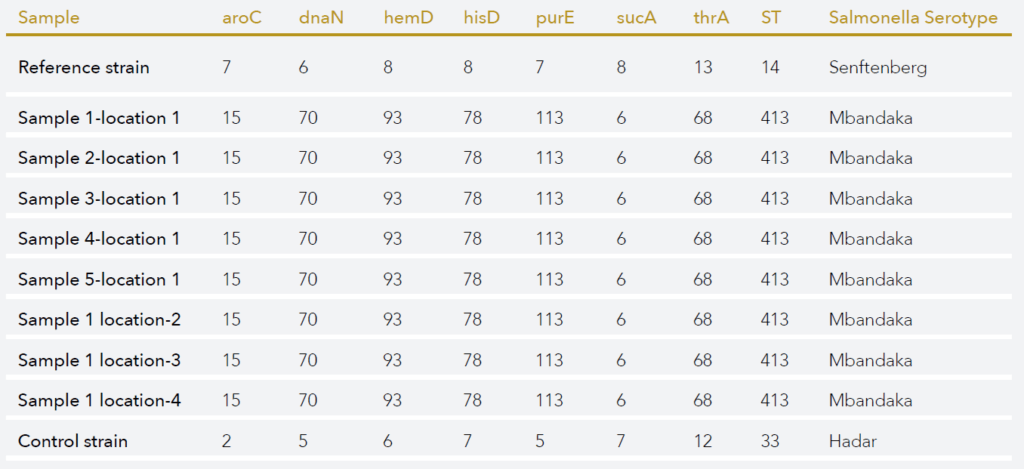
Whole genome sequencing (WGS) of microbial contaminants has become an effective method for investigating the information contained in the genome sequence. In addition, its highly discriminative power enables the comparison of genetic relatedness between bacteria and fungi even on a sub-species level. For this reason, WGS is being implemented worldwide and across sectors (pharma, biotech, food, veterinary and environment) for root cause analysis, the investigation of outbreaks, source attribution, and improved risk characterization models. In order to extract relevant information from the large quantity and complex data produced by WGS, BaseClear developed bioinformatics tools, allowing to analyse and interpret the data, starting from simple gene-searches to complex phylogenetic studies.